
Designing Biology: My Octant Software Engineering Internship
Designing Biology: My Octant Software Engineering Internship
This summer, I had the privilege to be a software engineering intern at Octant, working with a multidisciplinary team of scientists to help build a platform for the next generation of drug discovery. Along the way, I got a peek at some truly fascinating technology, and had plenty of fun too! I’m feeling more than ever before that biotech is the future.
Octant is an equal opportunity company that deeply values diversity, equity, and inclusion, and welcomes applicants of all backgrounds. We’re committed to growing the company with a diversity of talented individuals and our team is dedicated to fostering an inclusive and supportive work environment for everyone.

To solve this problem, I was tasked with creating a custom webapp for scientists to design experiments. Doing so would introduce a few major benefits:
- We could automate much of the complex spreadsheet functionality that scientists had to manually input.
- We could implement comprehensive data validation to notify users immediately about input errors, instead of them losing their work to a downstream software error after completing multiple additional steps.
- Scientists would no longer have to download and update software packages to run the scripts required to parse and manipulate the experimental planning data.
- Most importantly, a single URL would take any employee to the most recent stable version of the site. Whenever someone added a useful new feature, anyone could use it without having to dig through dozens of spreadsheets to find the most recent template.
Much of the infrastructure for what I needed had already been built. When I arrived, there was already a Python package called pyoctant for interacting with our lab instruments, and a webapp for storing experimental data (the "Platform Webapp"). Unfortunately, each of these projects had been designed by different teams to meet different needs, so they didn't integrate well with each other. After some discussion with my tech lead, Nicholas, we decided that the best way to integrate the various existing tools with each other would be to merge them into a single repository. We chose to use the existing Platform Webapp because it already had a database containing much of the data we would need to access, and because it used the Django framework, which would fit the needs of our new app.
Thus, the plan to extend the functionality of the Platform Webapp was born. Before writing any code, I began by interviewing scientists from different teams around the company to learn about the deficiencies of the old workflow, and what features would be required in the new app. In my conversations with scientists, the most common complaint I heard was that they would get all the way to the end of designing an experiment, and a script would crash because of a small mistake, sometimes leaving them clueless as to what had gone wrong. This problem required two complementary solutions: stringent validation would immediately identify user errors, and increased developer tooling would prevent the frequent software bugs that had plagued previous tools.
After making sure I understood the nuances of the problem, I worked closely with Nicholas to hammer out the data model and user flow for our site. Finally, I was ready to start writing code. I refactored the old app to have additional modules for the new features. Next, I containerized the database so that it could be run locally, and added typing and tests. All of this work was intended to help developers write more reliable code that would engender trust with the scientists.
With that work out of the way, I was ready to create the user interface. I studied up on the Django framework, and began implementing the website, page by page. With each page, I became more proficient and was able to design more complex forms, working my way from the first form, where a scientist can name an experiment and specify some basic metadata, to the final form, where scientists can edit the contents of individual wells that will hold chemicals.
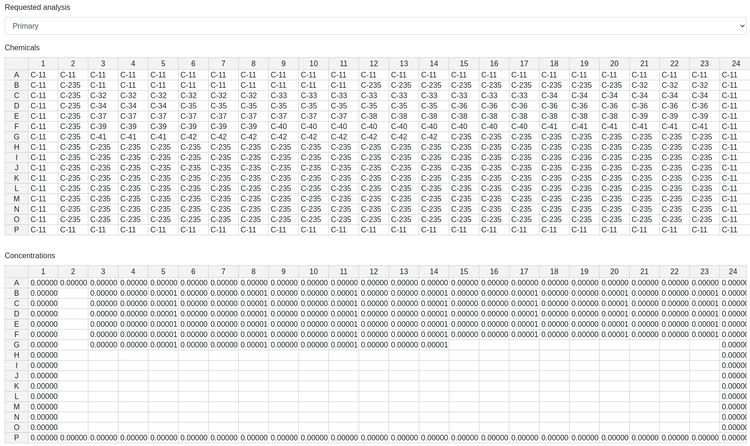

After a final code sprint, I was ready to present our new site to the company at an All Hands meeting. Although I was incredibly nervous, the demo went well and the scientists were excited to get their hands on the new tool!
Looking back on the summer, I learned an incredible amount about the practice of software engineering, from CI/CD to having conversations with users, and I learned even more about what it means to work in industry. Being lucky enough to have an in person internship as the Bay Area recovered from the pandemic, I had the pleasure of working closely with a talented, passionate group of scientists, engineers, and informaticians on problems that matter. I can't wait to see what Octant does next, and I'm counting the days until the first drug is approved that was discovered with the help of our new app.
Posted by

