
Octopus: A Simple, Low-Cost, and Scalable Plasmid Sequencing Method
Octopus: A Simple, Low-Cost, and Scalable Plasmid Sequencing Method
At Octant, we’re passionate about using the latest advances in synthetic biology to more rationally design drugs. In pursuit of our mission we developed OCTOPUS, a light-weight, cost-effective, and robust method for full-plasmid sequence verification using next-generation sequencing. OCTOPUS has accelerated much of Octant’s work over the last six months, and we think it will be broadly useful to the burgeoning community of startups and academics working to engineer biology.
Next-generation sequencing has fundamentally changed nearly every aspect of biological research. However, for the average scientist, it has yet to change the process of molecular cloning and sequence verification - a fundamental step in most molecular and synthetic biology R&D. Without access to expensive equipment and specialized processes, most cloning and sequence verification still uses outsourced Sanger Sequencing - a 60 year old technology that, while convenient at times, is quite painful at scale.
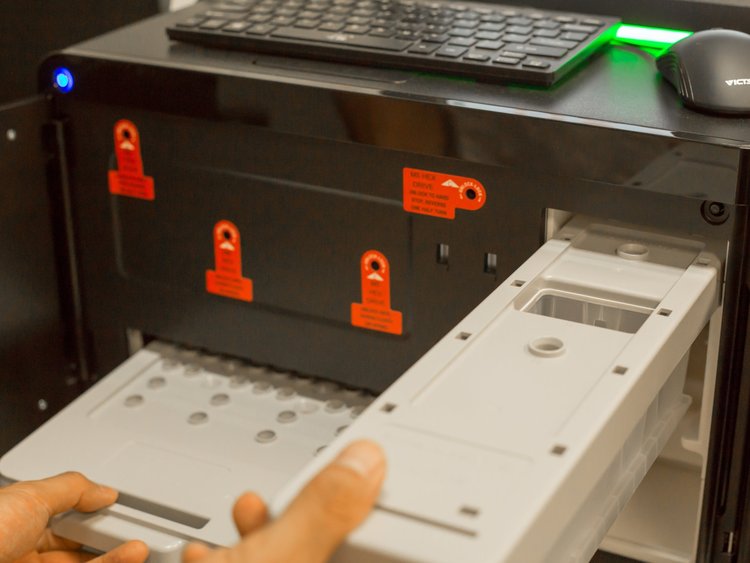

At Octant, we use a pooled cloning strategy that can generate hundreds of unique clones in a single reaction. Last year, as we scaled our cloning, we quickly outgrew our sequencing provider’s guaranteed turn-around time. They could commit to 192 individual reads per business day, which meant if we wanted to sequence 1000 colonies with four reads each, we could be waiting weeks for results. There were other limits to the existing Sanger-based process as well. For example, manually analyzing the data was about as painful as waiting for the results. In addition, we had to limit ourselves to only sequencing the parts of the plasmid we were changing. This is not ideal because it doesn’t find errors and mutations elsewhere in the plasmid, which when cloning at this scale, happen more than one would think. It became clear that our needs had outgrown the traditional approach. We needed a robust, scalable, and low touch method for verifying full plasmid sequences - ideally one that would not require a huge investment in labor and capital.
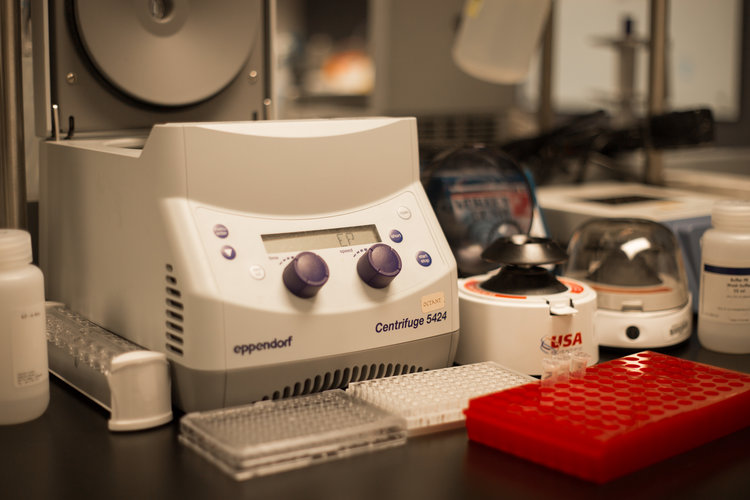

Around this time, we learned about Riptide, a new bacterial genome sequencing kit from iGenomX. In experimenting with Riptide we discovered that it works with crude lysate as the input. This combination is ideal for plasmid sequencing because it obviates the need for labor-intensive plasmid purification. We then built a turn-key computational pipeline to analyze the resulting data for plasmid verification - and thus OCTOPUS was born. OCTOPUS does not require specialized equipment or expensive liquid handling instruments, and the pipeline works on any modern desktop. All of this enables a single researcher to sequence over a thousand plasmids in less than a week.
-01.png)
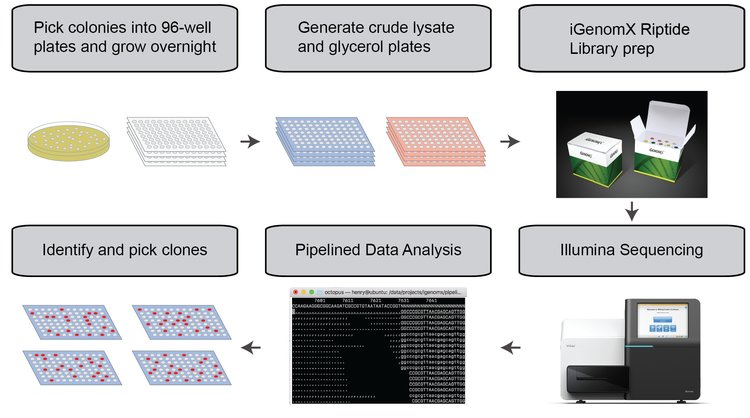
Here’s how OCTOPUS works in broad strokes:
- First, we pick colonies into normal 96-well plates and grow them overnight. Because we usually need 400-1200 colonies, it’s still easier and faster to do this manually than by colony picker. Also, colony pickers typically cost >$100K and don’t seem appropriate for our scale.
- We then take a small aliquot of the cultures, gently lyse the cells to preferentially release plasmid over genomic DNA, and save the remainder as glycerol stocks.
- Next, we input the crude lysate directly into the iGenomX Riptide kit at 1/2x scale. The kit provides even coverage of our plasmids, works in lysate, does not require low-volume acoustic handling, and is cost-effective. iGenomX has also been super supportive, responsive, and helpful in the development of OCTOPUS.
- Then, we sequence on a Miseq. We’ve found that sequencing 384 colonies with a V2 Micro MiSeq kit seems to be the sweet spot in terms of price per well and turn around time. If you want to scale up, we run three 384 well plates on a standard V2 kit. You could also sequence 96 wells with a nano kit, but the price per well will go up.
- We then feed the sequencing data to our analysis pipeline, which automatically performs a number of quality control and processing steps to analyze what’s in each well.
- We’re now ready to pick clones for plasmid purification. Since our pipeline provides unique plate-well identifiers, it’s simple to integrate the output into your liquid handler of choice for downstream processing (we use the OpenTron’s OT2).
You can find details of the full protocol and the software on our GitHub. We hope people will use and extend OCTOPUS to fit their own needs, share their improvements with the community, and provide feedback. More broadly, we hope that others, especially the hundreds of new startups in synthetic biology, consider sharing pre-competitive ideas, methods, tools, etc. We think the community will be stronger as a result.

OCTOPUS is the result of practicing a core value of Octant’s scientific culture - a tight integration between wet and digital experimentation enhances both. For example, we add molecular barcodes to the backbone of our plasmids (an optional feature of OCTOPUS), which greatly simplifies the computational detection of contaminants. Conversely, this feature makes the analysis pipeline robust to dirtier data, allowing the bench to take experimental shortcuts. Everything we do at Octant depends on this type of practical convergence, which leads to dramatic increases in productivity. If this kind of interplay is something that you geek out on, please consider joining us.
Posted by

Henry Chan

